Repository: systemml Updated Branches: refs/heads/master c3fdbb4da -> bda61b600
[MINOR] Updated the Linear Regression demo notebook Project: http://git-wip-us.apache.org/repos/asf/systemml/repo Commit: http://git-wip-us.apache.org/repos/asf/systemml/commit/bda61b60 Tree: http://git-wip-us.apache.org/repos/asf/systemml/tree/bda61b60 Diff: http://git-wip-us.apache.org/repos/asf/systemml/diff/bda61b60 Branch: refs/heads/master Commit: bda61b600a05e71be84848377b3e9ae93811c4d4 Parents: c3fdbb4 Author: Niketan Pansare <[email protected]> Authored: Fri Dec 7 15:31:48 2018 -0800 Committer: Niketan Pansare <[email protected]> Committed: Fri Dec 7 15:31:48 2018 -0800 ---------------------------------------------------------------------- .../Linear Regression Algorithms Demo.ipynb | 595 ------------------- .../Linear_Regression_Algorithms_Demo.ipynb | 582 ++++++++++++++++++ 2 files changed, 582 insertions(+), 595 deletions(-) ---------------------------------------------------------------------- http://git-wip-us.apache.org/repos/asf/systemml/blob/bda61b60/samples/jupyter-notebooks/Linear Regression Algorithms Demo.ipynb ---------------------------------------------------------------------- diff --git a/samples/jupyter-notebooks/Linear Regression Algorithms Demo.ipynb b/samples/jupyter-notebooks/Linear Regression Algorithms Demo.ipynb deleted file mode 100644 index 001f402..0000000 --- a/samples/jupyter-notebooks/Linear Regression Algorithms Demo.ipynb +++ /dev/null @@ -1,595 +0,0 @@ -{ - "cells": [ - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Linear Regression Algorithms using Apache SystemML\n", - "\n", - "This notebook shows:\n", - "- Install SystemML Python package and jar file\n", - " - pip\n", - " - SystemML 'Hello World'\n", - "- Example 1: Matrix Multiplication\n", - " - SystemML script to generate a random matrix, perform matrix multiplication, and compute the sum of the output\n", - " - Examine execution plans, and increase data size to obverve changed execution plans\n", - "- Load diabetes dataset from scikit-learn\n", - "- Example 2: Implement three different algorithms to train linear regression model\n", - " - Algorithm 1: Linear Regression - Direct Solve (no regularization)\n", - " - Algorithm 2: Linear Regression - Batch Gradient Descent (no regularization)\n", - " - Algorithm 3: Linear Regression - Conjugate Gradient (no regularization)\n", - "- Example 3: Invoke existing SystemML algorithm script LinearRegDS.dml using MLContext API\n", - "- Example 4: Invoke existing SystemML algorithm using scikit-learn/SparkML pipeline like API\n", - "- Uninstall/Clean up SystemML Python package and jar file" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Install SystemML Python package and jar file" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "!pip uninstall systemml --y\n", - "!pip install --user https://repository.apache.org/content/groups/snapshots/org/apache/systemml/systemml/1.0.0-SNAPSHOT/systemml-1.0.0-20171201.070207-23-python.tar.gz"; - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "!pip show systemml" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "### Import SystemML API " - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "from systemml import MLContext, dml, dmlFromResource\n", - "\n", - "ml = MLContext(sc)\n", - "\n", - "print \"Spark Version:\", sc.version\n", - "print \"SystemML Version:\", ml.version()\n", - "print \"SystemML Built-Time:\", ml.buildTime()" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "ml.execute(dml(\"\"\"s = 'Hello World!'\"\"\").output(\"s\")).get(\"s\")" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "### Import numpy, sklearn, and define some helper functions" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": true - }, - "outputs": [], - "source": [ - "import matplotlib.pyplot as plt\n", - "import numpy as np\n", - "from sklearn import datasets\n", - "plt.switch_backend('agg')" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Example 1: Matrix Multiplication" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "### SystemML script to generate a random matrix, perform matrix multiplication, and compute the sum of the output" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": true, - "slideshow": { - "slide_type": "-" - } - }, - "outputs": [], - "source": [ - "script = \"\"\"\n", - " X = rand(rows=$nr, cols=1000, sparsity=0.5)\n", - " A = t(X) %*% X\n", - " s = sum(A)\n", - "\"\"\"" - ] - }, - { - "cell_type": "raw", - "metadata": {}, - "source": [ - "ml.setStatistics(False)" - ] - }, - { - "cell_type": "raw", - "metadata": {}, - "source": [ - "ml.setExplain(True).setExplainLevel(\"runtime\")" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "prog = dml(script).input('$nr', 1e5).output('s')\n", - "s = ml.execute(prog).get('s')\n", - "print (s)" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Load diabetes dataset from scikit-learn " - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": true - }, - "outputs": [], - "source": [ - "%matplotlib inline" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "diabetes = datasets.load_diabetes()\n", - "diabetes_X = diabetes.data[:, np.newaxis, 2]\n", - "diabetes_X_train = diabetes_X[:-20]\n", - "diabetes_X_test = diabetes_X[-20:]\n", - "diabetes_y_train = diabetes.target[:-20].reshape(-1,1)\n", - "diabetes_y_test = diabetes.target[-20:].reshape(-1,1)\n", - "\n", - "plt.scatter(diabetes_X_train, diabetes_y_train, color='black')\n", - "plt.scatter(diabetes_X_test, diabetes_y_test, color='red')" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "diabetes.data.shape" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Example 2: Implement three different algorithms to train linear regression model" - ] - }, - { - "cell_type": "markdown", - "metadata": { - "collapsed": true - }, - "source": [ - "## Algorithm 1: Linear Regression - Direct Solve (no regularization) " - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "#### Least squares formulation\n", - "w* = argminw ||Xw-y||2 = argminw (y - Xw)'(y - Xw) = argminw (w'(X'X)w - w'(X'y))/2\n", - "\n", - "#### Setting the gradient\n", - "dw = (X'X)w - (X'y) to 0, w = (X'X)-1(X' y) = solve(X'X, X'y)" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": true - }, - "outputs": [], - "source": [ - "script = \"\"\"\n", - " # add constant feature to X to model intercept\n", - " X = cbind(X, matrix(1, rows=nrow(X), cols=1))\n", - " A = t(X) %*% X\n", - " b = t(X) %*% y\n", - " w = solve(A, b)\n", - " bias = as.scalar(w[nrow(w),1])\n", - " w = w[1:nrow(w)-1,]\n", - "\"\"\"" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "prog = dml(script).input(X=diabetes_X_train, y=diabetes_y_train).output('w', 'bias')\n", - "w, bias = ml.execute(prog).get('w','bias')\n", - "w = w.toNumPy()" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "plt.scatter(diabetes_X_train, diabetes_y_train, color='black')\n", - "plt.scatter(diabetes_X_test, diabetes_y_test, color='red')\n", - "\n", - "plt.plot(diabetes_X_test, (w*diabetes_X_test)+bias, color='blue', linestyle ='dotted')" - ] - }, - { - "cell_type": "markdown", - "metadata": { - "collapsed": true - }, - "source": [ - "## Algorithm 2: Linear Regression - Batch Gradient Descent (no regularization)" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "#### Algorithm\n", - "`Step 1: Start with an initial point \n", - "while(not converged) { \n", - " Step 2: Compute gradient dw. \n", - " Step 3: Compute stepsize alpha. \n", - " Step 4: Update: wnew = wold + alpha*dw \n", - "}`\n", - "\n", - "#### Gradient formula\n", - "`dw = r = (X'X)w - (X'y)`\n", - "\n", - "#### Step size formula\n", - "`Find number alpha to minimize f(w + alpha*r) \n", - "alpha = -(r'r)/(r'X'Xr)`\n", - "\n", - "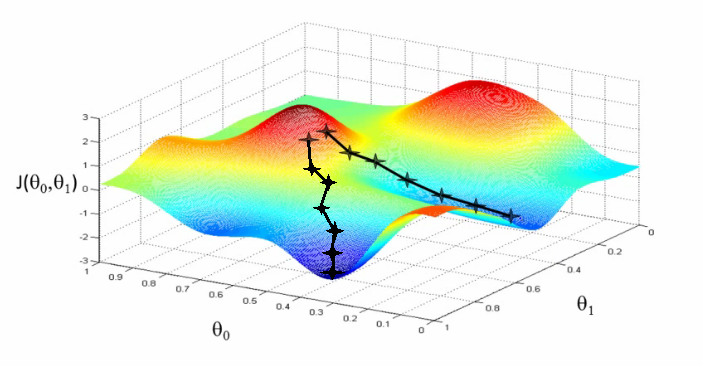" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": true - }, - "outputs": [], - "source": [ - "script = \"\"\"\n", - " # add constant feature to X to model intercepts\n", - " X = cbind(X, matrix(1, rows=nrow(X), cols=1))\n", - " max_iter = 100\n", - " w = matrix(0, rows=ncol(X), cols=1)\n", - " for(i in 1:max_iter){\n", - " XtX = t(X) %*% X\n", - " dw = XtX %*%w - t(X) %*% y\n", - " alpha = -(t(dw) %*% dw) / (t(dw) %*% XtX %*% dw)\n", - " w = w + dw*alpha\n", - " }\n", - " bias = as.scalar(w[nrow(w),1])\n", - " w = w[1:nrow(w)-1,] \n", - "\"\"\"" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "prog = dml(script).input(X=diabetes_X_train, y=diabetes_y_train).output('w', 'bias')\n", - "w, bias = ml.execute(prog).get('w', 'bias')\n", - "w = w.toNumPy()" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "plt.scatter(diabetes_X_train, diabetes_y_train, color='black')\n", - "plt.scatter(diabetes_X_test, diabetes_y_test, color='red')\n", - "\n", - "plt.plot(diabetes_X_test, (w*diabetes_X_test)+bias, color='red', linestyle ='dashed')" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Algorithm 3: Linear Regression - Conjugate Gradient (no regularization)" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Problem with gradient descent: Takes very similar directions many times\n", - "\n", - "Solution: Enforce conjugacy\n", - "\n", - "`Step 1: Start with an initial point \n", - "while(not converged) {\n", - " Step 2: Compute gradient dw.\n", - " Step 3: Compute stepsize alpha.\n", - " Step 4: Compute next direction p by enforcing conjugacy with previous direction.\n", - " Step 4: Update: w_new = w_old + alpha*p\n", - "}`\n", - "\n", - "\n" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": true - }, - "outputs": [], - "source": [ - "script = \"\"\"\n", - " # add constant feature to X to model intercepts\n", - " X = cbind(X, matrix(1, rows=nrow(X), cols=1))\n", - " m = ncol(X); i = 1; \n", - " max_iter = 20;\n", - " w = matrix (0, rows = m, cols = 1); # initialize weights to 0\n", - " dw = - t(X) %*% y; p = - dw; # dw = (X'X)w - (X'y)\n", - " norm_r2 = sum (dw ^ 2); \n", - " for(i in 1:max_iter) {\n", - " q = t(X) %*% (X %*% p)\n", - " alpha = norm_r2 / sum (p * q); # Minimizes f(w - alpha*r)\n", - " w = w + alpha * p; # update weights\n", - " dw = dw + alpha * q; \n", - " old_norm_r2 = norm_r2; norm_r2 = sum (dw ^ 2);\n", - " p = -dw + (norm_r2 / old_norm_r2) * p; # next direction - conjugacy to previous direction\n", - " i = i + 1;\n", - " }\n", - " bias = as.scalar(w[nrow(w),1])\n", - " w = w[1:nrow(w)-1,] \n", - "\"\"\"" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "prog = dml(script).input(X=diabetes_X_train, y=diabetes_y_train).output('w', 'bias')\n", - "w, bias = ml.execute(prog).get('w','bias')\n", - "w = w.toNumPy()" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "plt.scatter(diabetes_X_train, diabetes_y_train, color='black')\n", - "plt.scatter(diabetes_X_test, diabetes_y_test, color='red')\n", - "\n", - "plt.plot(diabetes_X_test, (w*diabetes_X_test)+bias, color='red', linestyle ='dashed')" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Example 3: Invoke existing SystemML algorithm script LinearRegDS.dml using MLContext API" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "prog = dmlFromResource('scripts/algorithms/LinearRegDS.dml').input(X=diabetes_X_train, y=diabetes_y_train).input('$icpt',1.0).output('beta_out')\n", - "w = ml.execute(prog).get('beta_out')\n", - "w = w.toNumPy()\n", - "bias=w[1]" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "plt.scatter(diabetes_X_train, diabetes_y_train, color='black')\n", - "plt.scatter(diabetes_X_test, diabetes_y_test, color='red')\n", - "\n", - "plt.plot(diabetes_X_test, (w[0]*diabetes_X_test)+bias, color='red', linestyle ='dashed')" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Example 4: Invoke existing SystemML algorithm using scikit-learn/SparkML pipeline like API" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "*mllearn* API allows a Python programmer to invoke SystemML's algorithms using scikit-learn like API as well as Spark's MLPipeline API." - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": true - }, - "outputs": [], - "source": [ - "from pyspark.sql import SQLContext\n", - "from systemml.mllearn import LinearRegression\n", - "sqlCtx = SQLContext(sc)" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "regr = LinearRegression(sqlCtx)\n", - "# Train the model using the training sets\n", - "regr.fit(diabetes_X_train, diabetes_y_train)" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "predictions = regr.predict(diabetes_X_test)" - ] - }, - { - "cell_type": "code", - "execution_count": null, - "metadata": { - "collapsed": false - }, - "outputs": [], - "source": [ - "# Use the trained model to perform prediction\n", - "%matplotlib inline\n", - "plt.scatter(diabetes_X_train, diabetes_y_train, color='black')\n", - "plt.scatter(diabetes_X_test, diabetes_y_test, color='red')\n", - "\n", - "plt.plot(diabetes_X_test, predictions, color='black')" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Uninstall/Clean up SystemML Python package and jar file" - ] - }, - { - "cell_type": "raw", - "metadata": {}, - "source": [ - "!pip uninstall systemml --y" - ] - } - ], - "metadata": { - "kernelspec": { - "display_name": "Python 2", - "language": "python", - "name": "python2" - }, - "language_info": { - "codemirror_mode": { - "name": "ipython", - "version": 2 - }, - "file_extension": ".py", - "mimetype": "text/x-python", - "name": "python", - "nbconvert_exporter": "python", - "pygments_lexer": "ipython2", - "version": "2.7.11" - } - }, - "nbformat": 4, - "nbformat_minor": 1 -} http://git-wip-us.apache.org/repos/asf/systemml/blob/bda61b60/samples/jupyter-notebooks/Linear_Regression_Algorithms_Demo.ipynb ---------------------------------------------------------------------- diff --git a/samples/jupyter-notebooks/Linear_Regression_Algorithms_Demo.ipynb b/samples/jupyter-notebooks/Linear_Regression_Algorithms_Demo.ipynb new file mode 100644 index 0000000..9e6f2a5 --- /dev/null +++ b/samples/jupyter-notebooks/Linear_Regression_Algorithms_Demo.ipynb @@ -0,0 +1,582 @@ +{ + "cells": [ + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Linear Regression Algorithms using Apache SystemML\n", + "\n", + "Table of Content:\n", + "- [Install SystemML using pip](#bullet1)\n", + "- [Example 1: Implement a simple 'Hello World' program in SystemML](#bullet2)\n", + "- [Example 2: Matrix Multiplication](#bullet3)\n", + "- [Load diabetes dataset from scikit-learn for the example 3](#bullet4)\n", + "- Example 3: Implement three different algorithms to train linear regression model\n", + " - [Algorithm 1: Linear Regression - Direct Solve (no regularization)](#example3algo1)\n", + " - [Algorithm 2: Linear Regression - Batch Gradient Descent (no regularization)](#example3algo2)\n", + " - [Algorithm 3: Linear Regression - Conjugate Gradient (no regularization)](#example3algo3)\n", + "- [Example 4: Invoke existing SystemML algorithm script LinearRegDS.dml using MLContext API](#example4)\n", + "- [Example 5: Invoke existing SystemML algorithm using scikit-learn/SparkML pipeline like API](#example5)\n", + "- [Uninstall/Clean up SystemML Python package and jar file](#uninstall)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Install SystemML using pip <a class=\"anchor\" id=\"bullet1\"></a>\n", + "\n", + "For more details, please see the [install guide](http://systemml.apache.org/install-systemml.html)." + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "!pip install --upgrade --user systemml" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "!pip show systemml" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Example 1: Implement a simple 'Hello World' program in SystemML <a class=\"anchor\" id=\"bullet2\"></a>\n", + "\n", + "### First import the classes necessary to implement the 'Hello World' program.\n", + "\n", + "The MLContext API offers a programmatic interface for interacting with SystemML from Spark using languages such as Scala, Java, and Python. As a result, it offers a convenient way to interact with SystemML from the Spark Shell and from Notebooks such as Jupyter and Zeppelin. Please refer to [the documentation](http://apache.github.io/systemml/spark-mlcontext-programming-guide) for more detail on the MLContext API.\n", + "\n", + "As a sidenote, here are alternative ways by which you can invoke SystemML (not covered in this notebook): \n", + "- Command-line invocation using either [spark-submit](http://apache.github.io/systemml/spark-batch-mode.html) or [hadoop](http://apache.github.io/systemml/hadoop-batch-mode.html).\n", + "- Using the [JMLC API](http://apache.github.io/systemml/jmlc.html)." + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "from systemml import MLContext, dml, dmlFromResource\n", + "\n", + "ml = MLContext(sc)\n", + "\n", + "print(\"Spark Version:\", sc.version)\n", + "print(\"SystemML Version:\", ml.version())\n", + "print(\"SystemML Built-Time:\", ml.buildTime())" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "# Step 1: Write the DML script\n", + "script = \"\"\"\n", + "print(\"Hello World!\");\n", + "\"\"\"\n", + "\n", + "# Step 2: Create a Python DML object\n", + "script = dml(script)\n", + "\n", + "# Step 3: Execute it using MLContext API\n", + "ml.execute(script)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Now let's implement a slightly more complicated 'Hello World' program where we initialize a string variable to 'Hello World!' and print it using Python. Note: we first register the output variable in the dml object (in the step 2) and then fetch it after execution (in the step 3)." + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "# Step 1: Write the DML script\n", + "script = \"\"\"\n", + "s = \"Hello World!\";\n", + "\"\"\"\n", + "\n", + "# Step 2: Create a Python DML object\n", + "script = dml(script).output('s')\n", + "\n", + "# Step 3: Execute it using MLContext API\n", + "s = ml.execute(script).get('s')\n", + "print(s)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Example 2: Matrix Multiplication <a class=\"anchor\" id=\"bullet3\"></a>\n", + "\n", + "Let's write a script to generate a random matrix, perform matrix multiplication, and compute the sum of the output." + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": { + "slideshow": { + "slide_type": "-" + } + }, + "outputs": [], + "source": [ + "# Step 1: Write the DML script\n", + "script = \"\"\"\n", + " # The number of rows is passed externally by the user via 'nr'\n", + " X = rand(rows=nr, cols=1000, sparsity=0.5)\n", + " A = t(X) %*% X\n", + " s = sum(A)\n", + "\"\"\"\n", + "\n", + "# Step 2: Create a Python DML object\n", + "script = dml(script).input(nr=1e5).output('s')\n", + "\n", + "# Step 3: Execute it using MLContext API\n", + "s = ml.execute(script).get('s')\n", + "print(s)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Now, let's generate a random matrix in NumPy and pass it to SystemML." + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "import numpy as np\n", + "npMatrix = np.random.rand(1000, 1000)\n", + "\n", + "# Step 1: Write the DML script\n", + "script = \"\"\"\n", + " A = t(X) %*% X\n", + " s = sum(A)\n", + "\"\"\"\n", + "\n", + "# Step 2: Create a Python DML object\n", + "script = dml(script).input(X=npMatrix).output('s')\n", + "\n", + "# Step 3: Execute it using MLContext API\n", + "s = ml.execute(script).get('s')\n", + "print(s)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Load diabetes dataset from scikit-learn for the example 3 <a class=\"anchor\" id=\"bullet4\"></a>" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "import matplotlib.pyplot as plt\n", + "import numpy as np\n", + "from sklearn import datasets\n", + "plt.switch_backend('agg')" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "%matplotlib inline" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "diabetes = datasets.load_diabetes()\n", + "diabetes_X = diabetes.data[:, np.newaxis, 2]\n", + "diabetes_X_train = diabetes_X[:-20]\n", + "diabetes_X_test = diabetes_X[-20:]\n", + "diabetes_y_train = diabetes.target[:-20].reshape(-1,1)\n", + "diabetes_y_test = diabetes.target[-20:].reshape(-1,1)\n", + "\n", + "plt.scatter(diabetes_X_train, diabetes_y_train, color='black')\n", + "plt.scatter(diabetes_X_test, diabetes_y_test, color='red')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Example 3: Implement three different algorithms to train linear regression model\n", + "\n", + "Linear regression models the relationship between one numerical response variable and one or more explanatory (feature) variables by fitting a linear equation to observed data. The feature vectors are provided as a matrix $X$ an the observed response values are provided as a 1-column matrix $y$.\n", + "\n", + "A linear regression line has an equation of the form $y = Xw$." + ] + }, + { + "cell_type": "markdown", + "metadata": { + "collapsed": true + }, + "source": [ + "### Algorithm 1: Linear Regression - Direct Solve (no regularization) <a class=\"anchor\" id=\"example3algo1\"></a>\n", + "\n", + "#### Least squares formulation\n", + "\n", + "The [least squares method](https://en.wikipedia.org/wiki/Least_squares) calculates the best-fitting line for the observed data by minimizing the sum of the squares of the difference between the predicted response $Xw$ and the actual response $y$.\n", + " \n", + "$w^* = argmin_w ||Xw-y||^2 \\\\\n", + "\\;\\;\\; = argmin_w (y - Xw)'(y - Xw) \\\\\n", + "\\;\\;\\; = argmin_w \\dfrac{(w'(X'X)w - w'(X'y))}{2}$\n", + "\n", + "To find the optimal parameter $w$, we set the gradient $dw = (X'X)w - (X'y)$ to 0.\n", + "\n", + "$(X'X)w - (X'y) = 0 \\\\\n", + "w = (X'X)^{-1}(X' y) \\\\\n", + " \\;\\;= solve(X'X, X'y)$" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "# Step 1: Write the DML script\n", + "script = \"\"\"\n", + " # add constant feature to X to model intercept\n", + " X = cbind(X, matrix(1, rows=nrow(X), cols=1))\n", + " A = t(X) %*% X\n", + " b = t(X) %*% y\n", + " w = solve(A, b)\n", + " bias = as.scalar(w[nrow(w),1])\n", + " w = w[1:nrow(w)-1,]\n", + "\"\"\"\n", + "\n", + "# Step 2: Create a Python DML object\n", + "script = dml(script).input(X=diabetes_X_train, y=diabetes_y_train).output('w', 'bias')\n", + "\n", + "# Step 3: Execute it using MLContext API\n", + "w, bias = ml.execute(script).get('w','bias')\n", + "w = w.toNumPy()" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "plt.scatter(diabetes_X_train, diabetes_y_train, color='black')\n", + "plt.scatter(diabetes_X_test, diabetes_y_test, color='red')\n", + "\n", + "plt.plot(diabetes_X_test, (w*diabetes_X_test)+bias, color='blue', linestyle ='dotted')" + ] + }, + { + "cell_type": "markdown", + "metadata": { + "collapsed": true + }, + "source": [ + "### Algorithm 2: Linear Regression - Batch Gradient Descent (no regularization) <a class=\"anchor\" id=\"example3algo2\"></a>" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Algorithm\n", + "`Step 1: Start with an initial point \n", + "while(not converged) { \n", + " Step 2: Compute gradient dw. \n", + " Step 3: Compute stepsize alpha. \n", + " Step 4: Update: wnew = wold + alpha*dw \n", + "}`\n", + "\n", + "#### Gradient formula\n", + "`dw = r = (X'X)w - (X'y)`\n", + "\n", + "#### Step size formula\n", + "`Find number alpha to minimize f(w + alpha*r) \n", + "alpha = -(r'r)/(r'X'Xr)`\n", + "\n", + "" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "# Step 1: Write the DML script\n", + "script = \"\"\"\n", + " # add constant feature to X to model intercepts\n", + " X = cbind(X, matrix(1, rows=nrow(X), cols=1))\n", + " max_iter = 100\n", + " w = matrix(0, rows=ncol(X), cols=1)\n", + " for(i in 1:max_iter){\n", + " XtX = t(X) %*% X\n", + " dw = XtX %*%w - t(X) %*% y\n", + " alpha = -(t(dw) %*% dw) / (t(dw) %*% XtX %*% dw)\n", + " w = w + dw*alpha\n", + " }\n", + " bias = as.scalar(w[nrow(w),1])\n", + " w = w[1:nrow(w)-1,] \n", + "\"\"\"\n", + "\n", + "# Step 2: Create a Python DML object\n", + "script = dml(script).input(X=diabetes_X_train, y=diabetes_y_train).output('w', 'bias')\n", + "\n", + "# Step 3: Execute it using MLContext API\n", + "w, bias = ml.execute(script).get('w','bias')\n", + "w = w.toNumPy()" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "plt.scatter(diabetes_X_train, diabetes_y_train, color='black')\n", + "plt.scatter(diabetes_X_test, diabetes_y_test, color='red')\n", + "\n", + "plt.plot(diabetes_X_test, (w*diabetes_X_test)+bias, color='red', linestyle ='dashed')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Algorithm 3: Linear Regression - Conjugate Gradient (no regularization) <a class=\"anchor\" id=\"example3algo3\"></a>" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Problem with gradient descent: Takes very similar directions many times\n", + "\n", + "Solution: Enforce conjugacy\n", + "\n", + "`Step 1: Start with an initial point \n", + "while(not converged) {\n", + " Step 2: Compute gradient dw.\n", + " Step 3: Compute stepsize alpha.\n", + " Step 4: Compute next direction p by enforcing conjugacy with previous direction.\n", + " Step 4: Update: w_new = w_old + alpha*p\n", + "}`\n", + "\n", + "\n" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "# Step 1: Write the DML script\n", + "script = \"\"\"\n", + " # add constant feature to X to model intercepts\n", + " X = cbind(X, matrix(1, rows=nrow(X), cols=1))\n", + " m = ncol(X); i = 1; \n", + " max_iter = 20;\n", + " w = matrix (0, rows = m, cols = 1); # initialize weights to 0\n", + " dw = - t(X) %*% y; p = - dw; # dw = (X'X)w - (X'y)\n", + " norm_r2 = sum (dw ^ 2); \n", + " for(i in 1:max_iter) {\n", + " q = t(X) %*% (X %*% p)\n", + " alpha = norm_r2 / sum (p * q); # Minimizes f(w - alpha*r)\n", + " w = w + alpha * p; # update weights\n", + " dw = dw + alpha * q; \n", + " old_norm_r2 = norm_r2; norm_r2 = sum (dw ^ 2);\n", + " p = -dw + (norm_r2 / old_norm_r2) * p; # next direction - conjugacy to previous direction\n", + " i = i + 1;\n", + " }\n", + " bias = as.scalar(w[nrow(w),1])\n", + " w = w[1:nrow(w)-1,] \n", + "\"\"\"\n", + "\n", + "# Step 2: Create a Python DML object\n", + "script = dml(script).input(X=diabetes_X_train, y=diabetes_y_train).output('w', 'bias')\n", + "\n", + "# Step 3: Execute it using MLContext API\n", + "w, bias = ml.execute(script).get('w','bias')\n", + "w = w.toNumPy()" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "plt.scatter(diabetes_X_train, diabetes_y_train, color='black')\n", + "plt.scatter(diabetes_X_test, diabetes_y_test, color='red')\n", + "\n", + "plt.plot(diabetes_X_test, (w*diabetes_X_test)+bias, color='red', linestyle ='dashed')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Example 4: Invoke existing SystemML algorithm script LinearRegDS.dml using MLContext API <a class=\"anchor\" id=\"example4\"></a>\n", + "\n", + "SystemML ships with several [pre-implemented algorithms](https://github.com/apache/systemml/tree/master/scripts/algorithms) that can be invoked directly. Please refer to the [algorithm reference manual](http://apache.github.io/systemml/algorithms-reference.html) for usage." + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "# Step 1: No need to write a DML script here. But, keeping it as a placeholder for consistency :)\n", + "\n", + "# Step 2: Create a Python DML object\n", + "script = dmlFromResource('scripts/algorithms/LinearRegDS.dml')\n", + "script = script.input(X=diabetes_X_train, y=diabetes_y_train).input('$icpt',1.0).output('beta_out')\n", + "\n", + "# Step 3: Execute it using MLContext API\n", + "w = ml.execute(script).get('beta_out')\n", + "w = w.toNumPy()\n", + "bias = w[1]\n", + "w = w[0]" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "plt.scatter(diabetes_X_train, diabetes_y_train, color='black')\n", + "plt.scatter(diabetes_X_test, diabetes_y_test, color='red')\n", + "\n", + "plt.plot(diabetes_X_test, (w*diabetes_X_test)+bias, color='red', linestyle ='dashed')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Example 5: Invoke existing SystemML algorithm using scikit-learn/SparkML pipeline like API <a class=\"anchor\" id=\"example5\"></a>" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "*mllearn* API allows a Python programmer to invoke SystemML's algorithms using scikit-learn like API as well as Spark's MLPipeline API." + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "# Step 1: No need to write a DML script here. But, keeping it as a placeholder for consistency :)\n", + "\n", + "# Step 2: No need to create a Python DML object. But, keeping it as a placeholder for consistency :)\n", + "\n", + "# Step 3: Execute Linear Regression using the mllearn API\n", + "from systemml.mllearn import LinearRegression\n", + "regr = LinearRegression(spark)\n", + "# Train the model using the training sets\n", + "regr.fit(diabetes_X_train, diabetes_y_train)" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "predictions = regr.predict(diabetes_X_test)" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "# Use the trained model to perform prediction\n", + "%matplotlib inline\n", + "plt.scatter(diabetes_X_train, diabetes_y_train, color='black')\n", + "plt.scatter(diabetes_X_test, diabetes_y_test, color='red')\n", + "\n", + "plt.plot(diabetes_X_test, predictions, color='black')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Uninstall/Clean up SystemML Python package and jar file <a class=\"anchor\" id=\"uninstall\"></a>" + ] + }, + { + "cell_type": "raw", + "metadata": {}, + "source": [ + "!pip uninstall systemml --y" + ] + } + ], + "metadata": { + "kernelspec": { + "display_name": "Python 2", + "language": "python", + "name": "python2" + }, + "language_info": { + "codemirror_mode": { + "name": "ipython", + "version": 2 + }, + "file_extension": ".py", + "mimetype": "text/x-python", + "name": "python", + "nbconvert_exporter": "python", + "pygments_lexer": "ipython2", + "version": "2.7.15" + } + }, + "nbformat": 4, + "nbformat_minor": 1 +}
