This is an automated email from the ASF dual-hosted git repository.
xinrong pushed a commit to branch master
in repository https://gitbox.apache.org/repos/asf/spark.git
The following commit(s) were added to refs/heads/master by this push:
new 1d3ec69dfdf [SPARK-40307][PYTHON] Introduce Arrow-optimized Python UDFs
1d3ec69dfdf is described below
commit 1d3ec69dfdf3edb0d688fb5294f8a17cc8f5e7e9
Author: Xinrong Meng <xinr...@apache.org>
AuthorDate: Thu Jan 12 20:23:20 2023 +0800
[SPARK-40307][PYTHON] Introduce Arrow-optimized Python UDFs
### What changes were proposed in this pull request?
Introduce Arrow-optimized Python UDFs. Please refer to
[design](https://docs.google.com/document/d/e/2PACX-1vQxFyrMqFM3zhDhKlczrl9ONixk56cVXUwDXK0MMx4Vv2kH3oo-tWYoujhrGbCXTF78CSD2kZtnhnrQ/pub)
for design details and micro benchmarks.
There are two ways to enable/disable the Arrow optimization for Python UDFs:
- the Spark configuration `spark.sql.execution.pythonUDF.arrow.enabled`,
disabled by default.
- the `useArrow` parameter of the `udf` function, None by default.
The Spark configuration takes effect only when `useArrow` is None.
Otherwise, `useArrow` decides whether a specific user-defined function is
optimized by Arrow or not.
The reason why we introduce these two ways is to provide both a convenient,
per-Spark-session control and a finer-grained, per-UDF control of the Arrow
optimization for Python UDFs.
### Why are the changes needed?
Python user-defined function (UDF) enables users to run arbitrary code
against PySpark columns. It uses Pickle for (de)serialization and executes row
by row.
One major performance bottleneck of Python UDFs is (de)serialization, that
is, the data interchanging between the worker JVM and the spawned Python
subprocess which actually executes the UDF.
The PR proposes a better alternative to handle the (de)serialization:
Arrow, which is used in the (de)serialization of Pandas UDF already.
#### Benchmark
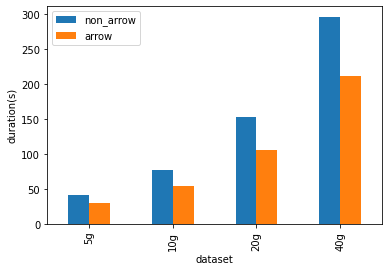
The micro benchmarks are conducted in a cluster with 1 driver (i3.2xlarge),
2 workers (i3.2xlarge). An i3.2xlarge machine has 61 GB Memory, 8 Cores. The
datasets used in the benchmarks are generated and sized 5 GB, 10 GB, 20 GB and
40 GB.
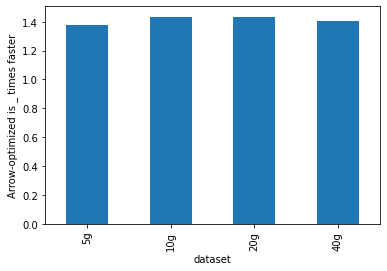
As shown below, Arrow-optimized Python UDFs are **~1.4x** faster than
non-Arrow-optimized Python UDFs.
Please refer to
[design](https://docs.google.com/document/d/e/2PACX-1vQxFyrMqFM3zhDhKlczrl9ONixk56cVXUwDXK0MMx4Vv2kH3oo-tWYoujhrGbCXTF78CSD2kZtnhnrQ/pub)
for details.
### Does this PR introduce _any_ user-facing change?
No, since the Arrow optimization for Python UDFs is disabled by default.
### How was this patch tested?
Unit tests.
Below is the script to generate the result table when the Arrow's type
coercion is needed, as in the
[docstring](https://github.com/apache/spark/pull/39384/files#diff-2df611ab00519d2d67e5fc20960bd5a6bd76ecd6f7d56cd50d8befd6ce30081bR96-R111)
of `_create_py_udf` .
```
import sys
import array
import datetime
from decimal import Decimal
from pyspark.sql import Row
from pyspark.sql.types import *
from pyspark.sql.functions import udf
data = [
None,
True,
1,
"a",
datetime.date(1970, 1, 1),
datetime.datetime(1970, 1, 1, 0, 0),
1.0,
array.array("i", [1]),
[1],
(1,),
bytearray([65, 66, 67]),
Decimal(1),
{"a": 1},
]
types = [
BooleanType(),
ByteType(),
ShortType(),
IntegerType(),
LongType(),
StringType(),
DateType(),
TimestampType(),
FloatType(),
DoubleType(),
BinaryType(),
DecimalType(10, 0),
]
df = spark.range(1)
results = []
count = 0
total = len(types) * len(data)
spark.sparkContext.setLogLevel("FATAL")
for t in types:
result = []
for v in data:
try:
row = df.select(udf(lambda _: v, t)("id")).first()
ret_str = repr(row[0])
except Exception:
ret_str = "X"
result.append(ret_str)
progress = "SQL Type: [%s]\n Python Value: [%s(%s)]\n Result
Python Value: [%s]" % (
t.simpleString(), str(v), type(v).__name__, ret_str)
count += 1
print("%s/%s:\n %s" % (count, total, progress))
results.append([t.simpleString()] + list(map(str, result)))
schema = ["SQL Type \\ Python Value(Type)"] + list(map(lambda v: "%s(%s)" %
(str(v), type(v).__name__), data))
strings = spark.createDataFrame(results, schema=schema)._jdf.showString(20,
20, False)
print("\n".join(map(lambda line: " # %s # noqa" % line,
strings.strip().split("\n"))))
```
Closes #39384 from xinrong-meng/arrow_py_udf_init.
Authored-by: Xinrong Meng <xinr...@apache.org>
Signed-off-by: Xinrong Meng <xinr...@apache.org>
---
dev/sparktestsupport/modules.py | 1 +
python/pyspark/sql/functions.py | 24 +++-
python/pyspark/sql/tests/test_arrow_python_udf.py | 130 +++++++++++++++++++++
python/pyspark/sql/tests/test_udf.py | 50 +++++++-
python/pyspark/sql/udf.py | 107 ++++++++++++++++-
.../org/apache/spark/sql/internal/SQLConf.scala | 9 ++
6 files changed, 315 insertions(+), 6 deletions(-)
diff --git a/dev/sparktestsupport/modules.py b/dev/sparktestsupport/modules.py
index 9d59680c5d2..9fa9430ba5f 100644
--- a/dev/sparktestsupport/modules.py
+++ b/dev/sparktestsupport/modules.py
@@ -467,6 +467,7 @@ pyspark_sql = Module(
"pyspark.sql.observation",
# unittests
"pyspark.sql.tests.test_arrow",
+ "pyspark.sql.tests.test_arrow_python_udf",
"pyspark.sql.tests.test_catalog",
"pyspark.sql.tests.test_column",
"pyspark.sql.tests.test_conf",
diff --git a/python/pyspark/sql/functions.py b/python/pyspark/sql/functions.py
index 60276b2a0b1..73698afa4e3 100644
--- a/python/pyspark/sql/functions.py
+++ b/python/pyspark/sql/functions.py
@@ -44,7 +44,7 @@ from pyspark.sql.dataframe import DataFrame
from pyspark.sql.types import ArrayType, DataType, StringType, StructType,
_from_numpy_type
# Keep UserDefinedFunction import for backwards compatible import; moved in
SPARK-22409
-from pyspark.sql.udf import UserDefinedFunction, _create_udf # noqa: F401
+from pyspark.sql.udf import UserDefinedFunction, _create_py_udf # noqa: F401
# Keep pandas_udf and PandasUDFType import for backwards compatible import;
moved in SPARK-28264
from pyspark.sql.pandas.functions import pandas_udf, PandasUDFType # noqa:
F401
@@ -9980,7 +9980,10 @@ def unwrap_udt(col: "ColumnOrName") -> Column:
@overload
def udf(
- f: Callable[..., Any], returnType: "DataTypeOrString" = StringType()
+ f: Callable[..., Any],
+ returnType: "DataTypeOrString" = StringType(),
+ *,
+ useArrow: Optional[bool] = None,
) -> "UserDefinedFunctionLike":
...
@@ -9988,6 +9991,8 @@ def udf(
@overload
def udf(
f: Optional["DataTypeOrString"] = None,
+ *,
+ useArrow: Optional[bool] = None,
) -> Callable[[Callable[..., Any]], "UserDefinedFunctionLike"]:
...
@@ -9996,6 +10001,7 @@ def udf(
def udf(
*,
returnType: "DataTypeOrString" = StringType(),
+ useArrow: Optional[bool] = None,
) -> Callable[[Callable[..., Any]], "UserDefinedFunctionLike"]:
...
@@ -10003,6 +10009,8 @@ def udf(
def udf(
f: Optional[Union[Callable[..., Any], "DataTypeOrString"]] = None,
returnType: "DataTypeOrString" = StringType(),
+ *,
+ useArrow: Optional[bool] = None,
) -> Union["UserDefinedFunctionLike", Callable[[Callable[..., Any]],
"UserDefinedFunctionLike"]]:
"""Creates a user defined function (UDF).
@@ -10015,6 +10023,9 @@ def udf(
returnType : :class:`pyspark.sql.types.DataType` or str
the return type of the user-defined function. The value can be either a
:class:`pyspark.sql.types.DataType` object or a DDL-formatted type
string.
+ useArrow : bool or None
+ whether to use Arrow to optimize the (de)serialization. When it is
None, the
+ Spark config "spark.sql.execution.pythonUDF.arrow.enabled" takes
effect.
Examples
--------
@@ -10093,10 +10104,15 @@ def udf(
# for decorator use it as a returnType
return_type = f or returnType
return functools.partial(
- _create_udf, returnType=return_type,
evalType=PythonEvalType.SQL_BATCHED_UDF
+ _create_py_udf,
+ returnType=return_type,
+ evalType=PythonEvalType.SQL_BATCHED_UDF,
+ useArrow=useArrow,
)
else:
- return _create_udf(f=f, returnType=returnType,
evalType=PythonEvalType.SQL_BATCHED_UDF)
+ return _create_py_udf(
+ f=f, returnType=returnType,
evalType=PythonEvalType.SQL_BATCHED_UDF, useArrow=useArrow
+ )
def _test() -> None:
diff --git a/python/pyspark/sql/tests/test_arrow_python_udf.py
b/python/pyspark/sql/tests/test_arrow_python_udf.py
new file mode 100644
index 00000000000..6b788d79848
--- /dev/null
+++ b/python/pyspark/sql/tests/test_arrow_python_udf.py
@@ -0,0 +1,130 @@
+#
+# Licensed to the Apache Software Foundation (ASF) under one or more
+# contributor license agreements. See the NOTICE file distributed with
+# this work for additional information regarding copyright ownership.
+# The ASF licenses this file to You under the Apache License, Version 2.0
+# (the "License"); you may not use this file except in compliance with
+# the License. You may obtain a copy of the License at
+#
+# http://www.apache.org/licenses/LICENSE-2.0
+#
+# Unless required by applicable law or agreed to in writing, software
+# distributed under the License is distributed on an "AS IS" BASIS,
+# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
+# See the License for the specific language governing permissions and
+# limitations under the License.
+#
+
+import unittest
+
+from pyspark.sql.functions import udf
+from pyspark.sql.tests.test_udf import BaseUDFTests
+from pyspark.testing.sqlutils import (
+ have_pandas,
+ have_pyarrow,
+ pandas_requirement_message,
+ pyarrow_requirement_message,
+ ReusedSQLTestCase,
+)
+
+
+@unittest.skipIf(
+ not have_pandas or not have_pyarrow, pandas_requirement_message or
pyarrow_requirement_message
+)
+class PythonUDFArrowTests(BaseUDFTests, ReusedSQLTestCase):
+ @classmethod
+ def setUpClass(cls):
+ super(PythonUDFArrowTests, cls).setUpClass()
+ cls.spark.conf.set("spark.sql.execution.pythonUDF.arrow.enabled",
"true")
+
+ @unittest.skip("Unrelated test, and it fails when it runs duplicatedly.")
+ def test_broadcast_in_udf(self):
+ super(PythonUDFArrowTests, self).test_broadcast_in_udf()
+
+ @unittest.skip("Unrelated test, and it fails when it runs duplicatedly.")
+ def test_register_java_function(self):
+ super(PythonUDFArrowTests, self).test_register_java_function()
+
+ @unittest.skip("Unrelated test, and it fails when it runs duplicatedly.")
+ def test_register_java_udaf(self):
+ super(PythonUDFArrowTests, self).test_register_java_udaf()
+
+ @unittest.skip("Struct input types are not supported with Arrow
optimization")
+ def test_udf_input_serialization_valuecompare_disabled(self):
+ super(PythonUDFArrowTests,
self).test_udf_input_serialization_valuecompare_disabled()
+
+ def test_nested_input_error(self):
+ with self.assertRaisesRegexp(
+ Exception, "NotImplementedError: Struct input type are not
supported"
+ ):
+ self.spark.range(1).selectExpr("struct(1, 2) as struct").select(
+ udf(lambda x: x)("struct")
+ ).collect()
+
+ def test_complex_input_types(self):
+ row = (
+ self.spark.range(1)
+ .selectExpr("array(1, 2, 3) as array", "map('a', 'b') as map")
+ .select(
+ udf(lambda x: str(x))("array"),
+ udf(lambda x: str(x))("map"),
+ )
+ .first()
+ )
+
+ # The input is NumPy array when the optimization is on.
+ self.assertEquals(row[0], "[1 2 3]")
+ self.assertEquals(row[1], "{'a': 'b'}")
+
+ def test_use_arrow(self):
+ # useArrow=True
+ row_true = (
+ self.spark.range(1)
+ .selectExpr(
+ "array(1, 2, 3) as array",
+ )
+ .select(
+ udf(lambda x: str(x), useArrow=True)("array"),
+ )
+ .first()
+ )
+
+ # useArrow=None
+ row_none = (
+ self.spark.range(1)
+ .selectExpr(
+ "array(1, 2, 3) as array",
+ )
+ .select(
+ udf(lambda x: str(x), useArrow=None)("array"),
+ )
+ .first()
+ )
+
+ # The input is a NumPy array when the Arrow optimization is on.
+ self.assertEquals(row_true[0], row_none[0]) # "[1 2 3]"
+
+ # useArrow=False
+ row_false = (
+ self.spark.range(1)
+ .selectExpr(
+ "array(1, 2, 3) as array",
+ )
+ .select(
+ udf(lambda x: str(x), useArrow=False)("array"),
+ )
+ .first()
+ )
+ self.assertEquals(row_false[0], "[1, 2, 3]")
+
+
+if __name__ == "__main__":
+ from pyspark.sql.tests.test_arrow_python_udf import * # noqa: F401
+
+ try:
+ import xmlrunner
+
+ testRunner = xmlrunner.XMLTestRunner(output="target/test-reports",
verbosity=2)
+ except ImportError:
+ testRunner = None
+ unittest.main(testRunner=testRunner, verbosity=2)
diff --git a/python/pyspark/sql/tests/test_udf.py
b/python/pyspark/sql/tests/test_udf.py
index 03bcbaf6ddf..fb669d158f5 100644
--- a/python/pyspark/sql/tests/test_udf.py
+++ b/python/pyspark/sql/tests/test_udf.py
@@ -43,7 +43,7 @@ from pyspark.testing.sqlutils import ReusedSQLTestCase,
test_compiled, test_not_
from pyspark.testing.utils import QuietTest
-class UDFTests(ReusedSQLTestCase):
+class BaseUDFTests(object):
def test_udf_with_callable(self):
d = [Row(number=i, squared=i**2) for i in range(10)]
rdd = self.sc.parallelize(d)
@@ -804,6 +804,54 @@ class UDFTests(ReusedSQLTestCase):
)
+class UDFTests(BaseUDFTests, ReusedSQLTestCase):
+ @classmethod
+ def setUpClass(cls):
+ super(BaseUDFTests, cls).setUpClass()
+ cls.spark.conf.set("spark.sql.execution.pythonUDF.arrow.enabled",
"false")
+
+
+def test_use_arrow(self):
+ # useArrow=True
+ row_true = (
+ self.spark.range(1)
+ .selectExpr(
+ "array(1, 2, 3) as array",
+ )
+ .select(
+ udf(lambda x: str(x), useArrow=True)("array"),
+ )
+ .first()
+ )
+ # The input is a NumPy array when the Arrow optimization is on.
+ self.assertEquals(row_true[0], "[1 2 3]")
+
+ # useArrow=None
+ row_none = (
+ self.spark.range(1)
+ .selectExpr(
+ "array(1, 2, 3) as array",
+ )
+ .select(
+ udf(lambda x: str(x), useArrow=None)("array"),
+ )
+ .first()
+ )
+
+ # useArrow=False
+ row_false = (
+ self.spark.range(1)
+ .selectExpr(
+ "array(1, 2, 3) as array",
+ )
+ .select(
+ udf(lambda x: str(x), useArrow=False)("array"),
+ )
+ .first()
+ )
+ self.assertEquals(row_false[0], row_none[0]) # "[1, 2, 3]"
+
+
class UDFInitializationTests(unittest.TestCase):
def tearDown(self):
if SparkSession._instantiatedSession is not None:
diff --git a/python/pyspark/sql/udf.py b/python/pyspark/sql/udf.py
index 7c7be392cd3..79ae456b1f7 100644
--- a/python/pyspark/sql/udf.py
+++ b/python/pyspark/sql/udf.py
@@ -17,6 +17,8 @@
"""
User-defined function related classes and functions
"""
+from inspect import getfullargspec
+
import functools
import inspect
import sys
@@ -30,12 +32,16 @@ from pyspark.profiler import Profiler
from pyspark.rdd import _prepare_for_python_RDD, PythonEvalType
from pyspark.sql.column import Column, _to_java_column, _to_seq
from pyspark.sql.types import (
- StringType,
+ ArrayType,
+ BinaryType,
DataType,
+ MapType,
+ StringType,
StructType,
_parse_datatype_string,
)
from pyspark.sql.pandas.types import to_arrow_type
+from pyspark.sql.pandas.utils import require_minimum_pandas_version,
require_minimum_pyarrow_version
if TYPE_CHECKING:
from pyspark.sql._typing import DataTypeOrString, ColumnOrName,
UserDefinedFunctionLike
@@ -75,6 +81,105 @@ def _create_udf(
return udf_obj._wrapped()
+def _create_py_udf(
+ f: Callable[..., Any],
+ returnType: "DataTypeOrString",
+ evalType: int,
+ useArrow: Optional[bool] = None,
+) -> "UserDefinedFunctionLike":
+ # The following table shows the results when the type coercion in Arrow is
needed, that is,
+ # when the user-specified return type(SQL Type) of the UDF and the actual
instance(Python
+ # Value(Type)) that the UDF returns are different.
+ # Arrow and Pickle have different type coercion rules, so a UDF might have
a different result
+ # with/without Arrow optimization. That's the main reason the Arrow
optimization for Python
+ # UDFs is disabled by default.
+ #
+-----------------------------+--------------+----------+------+---------------+--------------------+-----------------------------+----------+----------------------+---------+--------------------+----------------------------+------------+--------------+
# noqa
+ # |SQL Type \ Python Value(Type)|None(NoneType)|True(bool)|1(int)|
a(str)| 1970-01-01(date)|1970-01-01 00:00:00(datetime)|1.0(float)|array('i',
[1])(array)|[1](list)| (1,)(tuple)|bytearray(b'ABC')(bytearray)|
1(Decimal)|{'a': 1}(dict)| # noqa
+ #
+-----------------------------+--------------+----------+------+---------------+--------------------+-----------------------------+----------+----------------------+---------+--------------------+----------------------------+------------+--------------+
# noqa
+ # | boolean| None| True| None|
None| None| None| None|
None| None| None| None|
None| None| # noqa
+ # | tinyint| None| None| 1|
None| None| None| None|
None| None| None| None|
None| None| # noqa
+ # | smallint| None| None| 1|
None| None| None| None|
None| None| None| None|
None| None| # noqa
+ # | int| None| None| 1|
None| None| None| None|
None| None| None| None|
None| None| # noqa
+ # | bigint| None| None| 1|
None| None| None| None|
None| None| None| None|
None| None| # noqa
+ # | string| None| 'true'| '1'|
'a'|'java.util.Gregor...| 'java.util.Gregor...| '1.0'|
'[I@120d813a'| '[1]'|'[Ljava.lang.Obje...| '[B@48571878'|
'1'| '{a=1}'| # noqa
+ # | date| None| X| X|
X|datetime.date(197...| datetime.date(197...| X|
X| X| X| X|
X| X| # noqa
+ # | timestamp| None| X| X|
X| X| datetime.datetime...| X|
X| X| X| X|
X| X| # noqa
+ # | float| None| None| None|
None| None| None| 1.0|
None| None| None| None|
None| None| # noqa
+ # | double| None| None| None|
None| None| None| 1.0|
None| None| None| None|
None| None| # noqa
+ # | binary| None| None|
None|bytearray(b'a')| None| None|
None| None| None| None|
bytearray(b'ABC')| None| None| # noqa
+ # | decimal(10,0)| None| None| None|
None| None| None| None|
None| None| None|
None|Decimal('1')| None| # noqa
+ #
+-----------------------------+--------------+----------+------+---------------+--------------------+-----------------------------+----------+----------------------+---------+--------------------+----------------------------+------------+--------------+
# noqa
+ # Note: Python 3.9.15, Pandas 1.5.2 and PyArrow 10.0.1 are used.
+ # Note: The values of 'SQL Type' are DDL formatted strings, which can be
used as `returnType`s.
+ # Note: The values inside the table are generated by `repr`. X' means it
throws an exception
+ # during the conversion.
+
+ from pyspark.sql import SparkSession
+
+ session = SparkSession._instantiatedSession
+ if session is None:
+ is_arrow_enabled = False
+ else:
+ is_arrow_enabled = (
+ session.conf.get("spark.sql.execution.pythonUDF.arrow.enabled") ==
"true"
+ if useArrow is None
+ else useArrow
+ )
+
+ regular_udf = _create_udf(f, returnType, evalType)
+ return_type = regular_udf.returnType
+ try:
+ is_func_with_args = len(getfullargspec(f).args) > 0
+ except TypeError:
+ is_func_with_args = False
+ is_output_atomic_type = (
+ not isinstance(return_type, StructType)
+ and not isinstance(return_type, MapType)
+ and not isinstance(return_type, ArrayType)
+ )
+ if is_arrow_enabled and is_output_atomic_type and is_func_with_args:
+ require_minimum_pandas_version()
+ require_minimum_pyarrow_version()
+
+ import pandas as pd
+ from pyspark.sql.pandas.functions import _create_pandas_udf # type:
ignore[attr-defined]
+
+ # "result_func" ensures the result of a Python UDF to be consistent
with/without Arrow
+ # optimization.
+ # Otherwise, an Arrow-optimized Python UDF raises
"pyarrow.lib.ArrowTypeError: Expected a
+ # string or bytes dtype, got ..." whereas a non-Arrow-optimized Python
UDF returns
+ # successfully.
+ result_func = lambda pdf: pdf # noqa: E731
+ if type(return_type) == StringType:
+ result_func = lambda r: str(r) if r is not None else r # noqa:
E731
+ elif type(return_type) == BinaryType:
+ result_func = lambda r: bytes(r) if r is not None else r # noqa:
E731
+
+ def vectorized_udf(*args: pd.Series) -> pd.Series:
+ if any(map(lambda arg: isinstance(arg, pd.DataFrame), args)):
+ raise NotImplementedError(
+ "Struct input type are not supported with Arrow
optimization "
+ "enabled in Python UDFs. Disable "
+ "'spark.sql.execution.pythonUDF.arrow.enabled' to
workaround."
+ )
+ return pd.Series(result_func(f(*a)) for a in zip(*args))
+
+ # Regular UDFs can take callable instances too.
+ vectorized_udf.__name__ = f.__name__ if hasattr(f, "__name__") else
f.__class__.__name__
+ vectorized_udf.__module__ = (
+ f.__module__ if hasattr(f, "__module__") else
f.__class__.__module__
+ )
+ vectorized_udf.__doc__ = f.__doc__
+ pudf = _create_pandas_udf(vectorized_udf, returnType, None)
+ # Keep the attributes as if this is a regular Python UDF.
+ pudf.func = f
+ pudf.returnType = return_type
+ pudf.evalType = regular_udf.evalType
+ return pudf
+ else:
+ return regular_udf
+
+
class UserDefinedFunction:
"""
User defined function in Python
diff --git
a/sql/catalyst/src/main/scala/org/apache/spark/sql/internal/SQLConf.scala
b/sql/catalyst/src/main/scala/org/apache/spark/sql/internal/SQLConf.scala
index abde198617d..fa69ab0ce38 100644
--- a/sql/catalyst/src/main/scala/org/apache/spark/sql/internal/SQLConf.scala
+++ b/sql/catalyst/src/main/scala/org/apache/spark/sql/internal/SQLConf.scala
@@ -2761,6 +2761,15 @@ object SQLConf {
// show full stacktrace in tests but hide in production by default.
.createWithDefault(!Utils.isTesting)
+ val PYTHON_UDF_ARROW_ENABLED =
+ buildConf("spark.sql.execution.pythonUDF.arrow.enabled")
+ .doc("Enable Arrow optimization in regular Python UDFs. This
optimization " +
+ "can only be enabled for atomic output types and input types except
struct and map types " +
+ "when the given function takes at least one argument.")
+ .version("3.4.0")
+ .booleanConf
+ .createWithDefault(false)
+
val PANDAS_GROUPED_MAP_ASSIGN_COLUMNS_BY_NAME =
buildConf("spark.sql.legacy.execution.pandas.groupedMap.assignColumnsByName")
.internal()
---------------------------------------------------------------------
To unsubscribe, e-mail: commits-unsubscr...@spark.apache.org
For additional commands, e-mail: commits-h...@spark.apache.org
